Project Type: Regular FONDECYT #1190701
Position: Head Researcher
Award Year: 2019. Ending Year: 2021.

Description
The study of the human connectome is an interesting, growing and key research area. It has led to the creation of initiatives such as the Human Connectome Project (http://www.humanconnectomeproject.org/) and the Human Brain Project (https://www.humanbrainproject.eu/). Their objective is to fully describe the brain’s structure and connections, in order to better understand its functioning. This analysis involves different modalities, such as anatomical, functional, and clinical data, among others.
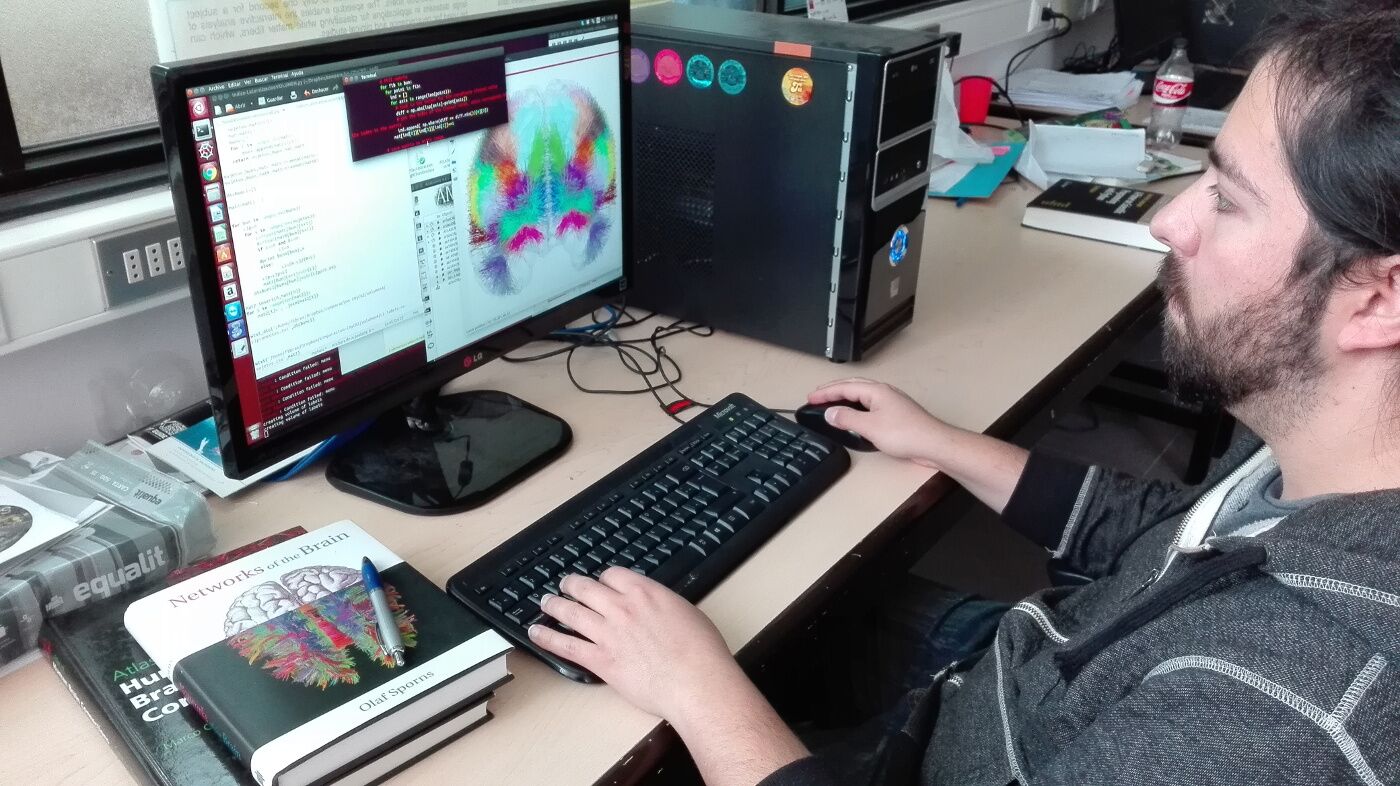
This ongoing project focuses on contributing to the decoding of the human brain connectome, via the development of methods for the analysis of brain connectivity based on high-quality diffusion Magnetic Resonance Imaging (dMRI) databases. It also aims to develop interactive tools for the analysis of WM bundles and structure, specifically regarding the visualization, segmentation, and manipulation of fiber tractography data.
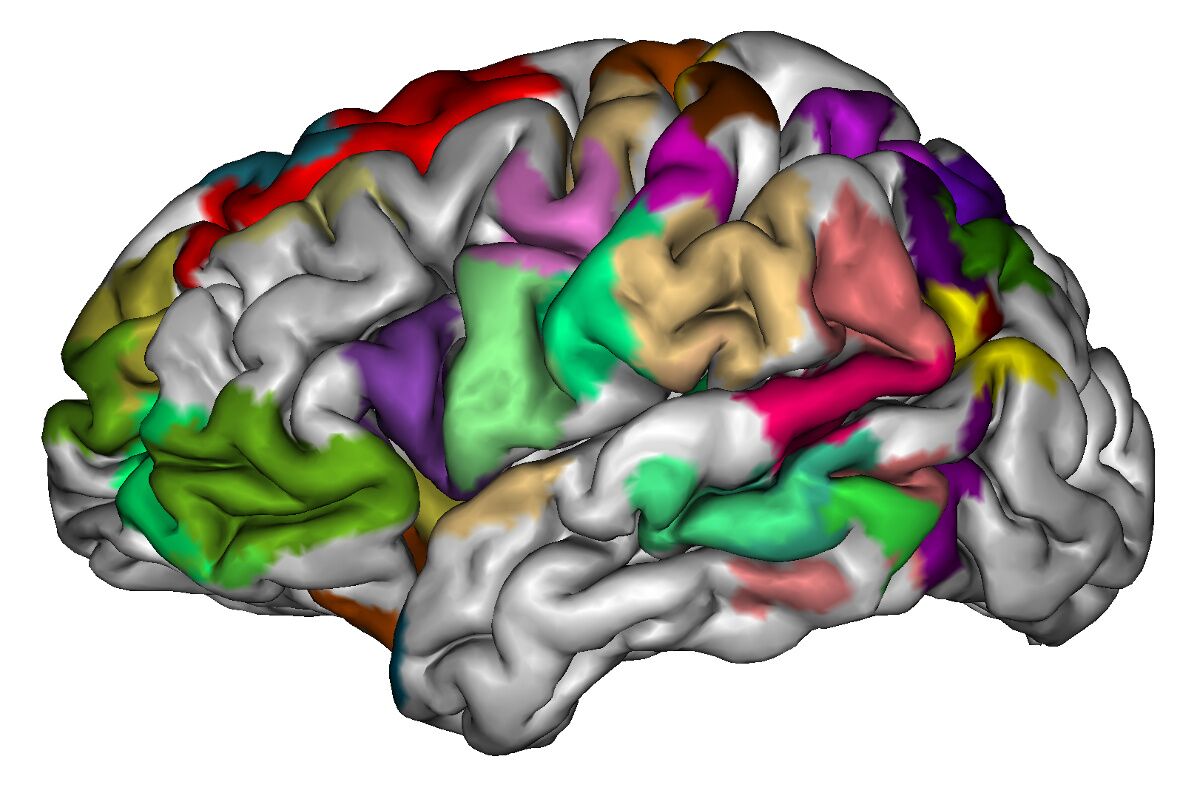
The main goal of this project is the development of new tractography-based cortical parcellation methods for a better description of the human brain connectome, using high-quality MRI data. These tools will not only help achieve the goal of describing the human connectome, but will contribute to the development of neuroimaging research in Chile as well. This will be of great interest for neuroscientists, neurologists, and neuroanatomist, as it will allow performing more accurate and detailed clinical studies, and thus also answer fundamental questions about brain structure and function.
The specific goals are:
Undergraduate Theses related to the project
| Student | Thesis | Defense Date |
|---|---|---|
| Paulo Olivares | Informatics Engineering Thesis (co-advised): “GPGPU implementation of a brain fiber clustering method based on point distribution” | May 11th, 2020. |
| Isaías Huerta | Informatics Engineering Thesis (co-advised): “Inter-subject clustering of brain fibers calculated from tractography” | January 29th, 2020. |
| Felipe Silva V. | Biomedical Engineering Thesis: “Cortical parcellation method based on a graph representation of brain connections” | April 1st, 2019. |
Postgraduate Theses related to the project
| Student | Thesis | Defense Date |
|---|---|---|
| Andrea Vásquez V. | Master in Computer Sciences. “Efficient brain fiber clustering method based on point distribution” |
April 16th, 2019. |